Statistics in EEGLAB
Computing statistics is essential to the observation of group, session, and/or condition measure differences. EEGLAB allows users to use either parametric or non-parametric statistics to compute and estimate the reliability of these differences across conditions and/or groups. Here we describe some essential concepts behind the statistical methods implemented in EEGLAB. For a complete introduction to robust statistics in EEG research, you may watch this series of short videos. Click on the icon on the top right corner to access the list of videos in the playlist.
Table of contents
Different types of inferential statistics
What is the best way of doing statistics on your data? Do you have continuous numerical data, categorical/discrete data? For discrete data, you may use binomial, chi2, or Cochran’s Q test, depending on your design. We will assume here that you have continuous numerical EEG data.
Parametric statistics
If your data is Gaussian, you will want to use t-test (paired or unpaired), repeated measure ANOVA if you have more than two conditions or n x m design. Note that depending on your design, some complex version of ANOVA might be required. For example, if you want to blend ANOVA and regression, you will have to use an ANCOVA. Note that even if your data is not Gaussian, parametric tests may be used. However, their power may be reduced compared to other tests.
Some transformation might be necessary in some cases to make the data more Gaussian. The most common is to log-transform spectral EEG power.
Non-parametric statistics
Equivalent non-parametric tests exist for all the parametric tests. These tests assume a Gaussian probability distribution of the rank of the sorted data values. (t-test paired -> Sign test or Wilcoxon test; unpaired t-test -> Mann Whitney U test; ANOVA -> Krukal Walis and Friedman test). These types of inferential statistics are usually not used on EEG data and are not available in EEGLAB.
Use of surrogate tests for non-Gaussian data
Surrogate tests are ideals for EEG data because they make no assumption on the data distribution. Surrogate tests consist of repetitively shuffling values between conditions and recompute the measure of interest using the shuffled data (for example, the difference between 2 conditions). We then obtain a distribution of difference, and we can see if the original difference is in the tail of this distribution. Under the null hypothesis of no difference between the conditions, the original difference should not be in the tail. If it is, we can assess the probability of rejecting H0. Permutation and bootstrap test are two surrogate tests. Permutation performs drawing of data samples without replacement, and bootstrap performs drawing with replacement. In theory, bootstrap is more valid since draws are independent of each other. In practice, there is little difference between the two tests. The main drawback of such approaches is that they can take longer to compute.
The cleaner the data, the easier the statistics. But getting clean data is an effort in itself. When outliers are present, surrogate tests are the most robust. However, even a surrogate test may fail in these conditions. The solution is to trim the distribution of value at each iteration of the test.
We recommend using parametric tests for data exploration since they are fast to compute. However, for publication, we recommend using surrogate tests.
Statistics implemented in EEGLAB
EEGLAB allows performing classical parametric tests (paired t-test, unpaired t-test, ANOVA) on ERPs, power spectra, ERSPs, and ITCs.
Below, we will use channel ERPs as an example, though in general, we recommend source-resolved measures be used instead. This is because no data features of interest are generated in the scalp, but rather in the brain itself.
For example, given 15 subjects’ ERPs for two task or stimulus conditions, EEGLAB functions can perform a simple two-tailed paired t-test at each trial latency on the average ERPs from each subject.
If there are different numbers of subjects in each condition, EEGLAB will use an unpaired t-test. If there are more than two STUDY conditions, EEGLAB will use ANOVA instead of a t-test. For mean power spectra, the p-values are computed at every frequency; for ERSP and ITC time/frequency transforms, p-values are computed at every time/frequency point.
EEGLAB functions can also compute non-parametric statistics. The null hypothesis is that there is no difference between the conditions. In this case, the average difference between the ERPs for two conditions should lie within the average difference between ‘surrogate’ grand mean condition ERPs, averages of ERPs from the two conditions whose condition assignments have been shuffled randomly. An example follows:
Given 15 subjects and two conditions, let us use a1, a2, a3, … a15, the scalp channel ERP values (in microvolts) at 300 ms for all 15 subjects in the first condition, and b1, b2, b3, … b15, the ERP values for the second condition.
The grand average ERP condition difference is
d = mean( (a1-b1) + (a2-b2) + … + (a15-b15) ).
Now, if we repeatedly shuffle these values between the two condition (under the null hypothesis that there are no significant differences between them), and then average the shuffled values,
d1 = mean( (b1-a1) + (a2-b2) + … + (b15-a15) ).
d2 = mean( (a1-b1) + (b2-a2) + … + (a15-b15) ).
d3 = mean( (b1-a1) + (b2-a2) + … + (a15-b15) ). …
We then obtain a distribution of surrogate condition-mean ERP values dx constructed using the null hypothesis (see their smoothed histogram below). If we observe that the initial value d lies in the very tail of this surrogate value distribution, then the supposed null hypothesis (no difference between conditions) may be rejected as highly unlikely, and the observed condition difference may be said to be statistically valid or significant.
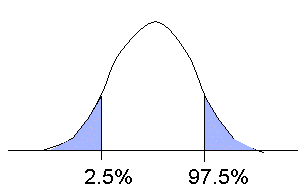
Note that the surrogate value distribution above can take any shape and does not need to be gaussian. In practice, we do not compute the mean condition ERP difference, but its t-value (the mean difference divided by the standard deviation of the difference and multiplied by the square root of the number of observations less one). The result is equivalent to using the mean difference. The advantage is that when we have more conditions, we can use the comparable ANOVA measure. Computing the probability density distribution of the t-test or ANOVA is only a “trick” to be able to obtain a distance measure across all subjects and conditions. It has nothing to do with relying on a parametric t-test or ANOVA model, which assume underlying gaussian value distributions.
Correcting for multiple comparisons
When performing a large number of statistical inferences, it is necessary to correct for multiple comparisons. For example, with a statistical threshold at p<0.05, by definition, about 5% of the inferred significant values will be false positives. We advise watching the short video on correction for multiple comparisons on Youtube.
There are several methods for correcting for multiple comparisons.
-
Bonferroni: The most conservative method, the Bonferroni method, simply divides the p-value by the number of comparisons. For example, when computing ERSP time-frequency images of 100 frequencies by 200 time points, the number of inferences is 20,000. To correct for multiple comparisons at p<0.05, a statistical threshold of 0.05/20000 = 0.0000025 should be applied. This method is quite conservative as, essentially, it assumes (erroneously) that all time/frequency values are independent.
-
Holm’s method: Holm’s method, also called Holm-Bonferroni’s method, is not as conservative. Actual uncorrected p-values are sorted and, to assess whether a given p-value reaches the corrected threshold for multiple comparisons, the lowest uncorrected p-value is compared to the Bonferroni statistical threshold of 0.05/20000. Next, the second-lowest is compared to a statistical threshold of 0.05/(20000-1), etc. The highest uncorrected p-value is compared to the uncorrected threshold of 0.05/(20000-19999)=0.05.
- False Discovery Rate: The False Discovery Rate (FDR) method corrects for the percentage of false positives (no more than 0.05% false positives with a 0.05 p-value threshold). This is different from Bonferroni and Holm-Bonferroni, which correct for the family-wise error rate (aiming to achieve no more false positives than when performing a single statistical test). FDR and Holm-Bonferroni use the same procedure to assess significance, except for FDR, the gradient of p-value threshold between the Bonferroni corrected, and uncorrected p-value is linear (while it is inverse for Holm-Bonferroni).
In EEGLAB, other methods are made available using statistics routines written for FieldTrip and LIMO – the max, cluster, and TFCE methods. These methods are now widely been used, but are only available when using non-parametric (surrogate data-based statistics).
-
Max method: At each iteration in computing a surrogate distribution of a time-frequency decomposition (for example), the maximum statistic across all time-frequency points is calculated. The surrogate distribution is compiled of these maximum statistics. The original statistics (for example, t-scores) at all time-frequency points are compared against this unique surrogate distribution (instead of each time-frequency point being compared to its corresponding surrogate distribution as in the other methods).
-
Cluster method: The cluster method is also only available when using non-parametric (surrogate) statistics. It is similar to the max method. Instead of using the raw statistics, it uses the size of significant regions (uncorrected) as the statistics.
-
Threshold free cluster enhancement (TFCE) method: The TFCE method is only available in the EEGLAB LIMO plugin. It consists in enhancing statistical values (for example, t-statistics) if they belong to a cluster of similar values. After cluster enhancement, it then uses the max method.
General Linear Modelling in EEGLAB
For complex design, you might also want to build a design matrix as this is done in the SPM software package for processing fMRI data, for example. Once you have design matrix, you may use a general linear model to fit parameters. The LIMO statistics video series introduces general linear modeling using EEGLAB and the LIMO toolbox. General linear models encompass all linear statistics and offer a general framework for performing statistics on EEG data.
Additional tips and resources
Do not: p-hacking consist in testing all possible combination of test and data transformation in the hope that one test will be significant (and it usually will!). Obviously this type of practice should be discouraged. In practice, we like to check that several inference tests and methods for correcting for multiple comparisons and report all results in articles.
We suggest consulting a relevant statistics book for more details: An introduction to statistics written by Arnaud Delorme is available here. We also recommend Rand Wilcox’s textbook on non-parametric statistics Introduction to robust estimation and hypothesis testing.