Batch processing of subjects’ data
In this tutorial, we introduce advanced elements of EEGLAB focused on processing multiple subjects. These topics are recommended to successfully analyze large-scale experiments, which go beyond single-subject analysis.
In addition to reading this tutorial, you may want to watch the short video below on multiple subjects processing in EEGLAB:
Table of contents
Load multiple datasets
To explore the multiple-dataset processing functionality of EEGLAB, you must first load and select several datasets.
In this tutorial, we will use a 5-subject experiment (450Mb). See the STUDY creation tutorial for more information on this data.
After uncompressing the data archive, load one by one all the datasets in EEGLAB using the File → Load existing dataset menu item. When several datasets are in the same folder, they may be all selected and loaded in EEGLAB simultaneously, as shown below.
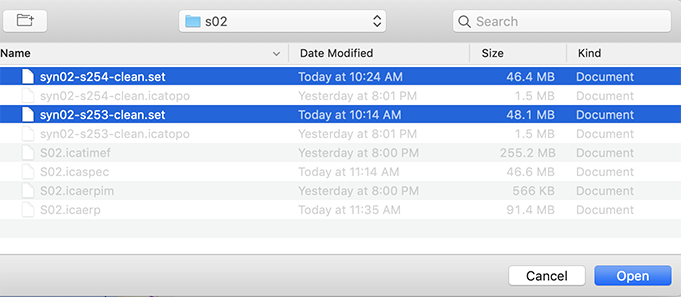
Select multiple datasets for processing
After importing all ten datasets, select the Datasets → Select multiple datasets menu item, as shown below.
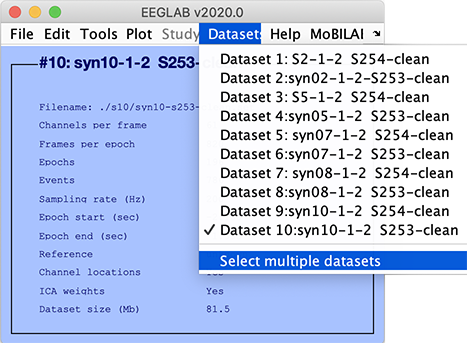
The following GUI will pop up. Select all ten datasets and press Ok.
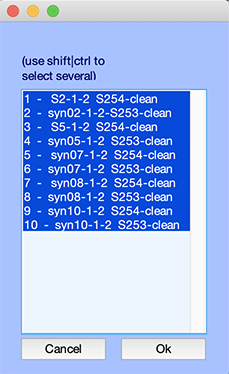
Perform batch processing
EEGLAB allows processing a collection of datasets, whether these datasets are organized in a STUDY or not. When there are multiple current datasets, menu items unable to process multiple datasets are disabled. All available tools process data in a similar way. Upon menu selection, a menu window pops up (identical to the single dataset window) in which you may select processing parameters that are then applied to all the datasets.
Note that the behavior of EEGLAB will differ depending on your optional settings under File → Preferences.
-
If you allow only one dataset to be present in memory at a time (see the dataset management section of the tutorial sfor more details), existing datasets will be automatically overwritten on disk (a warning window will appear).
-
However, suppose you allow all datasets to be present in memory simultaneously. In that case, only the datasets in memory will be overwritten, and their copies in disk files will not be affected (you may then select menu item File → Save current dataset(s) to save all the currently selected datasets).
You may now select any available menu item to process the selected datasets. A standard sequence to process raw datasets could be:
- Remove unwanted channels using the Edit → Select data menu item.
- Look up channel locations using the Edit → Channel locations menu item.
- High-pass filter the data using the Tools → Filter the data → Basic FIR filter menu item.
- Perform automated artifact rejection using the Tools → Reject data using Clean Rawdata and ASR menu item.
- Re-reference the data using the Tools → Re-reference the data menu item.
- Run independent component analysis using the Tools → Decompose data by ICA menu item.
- Label components using the Tools → Classify components using IClabel → Label components menu item.
- Classify components using the Tools → Classify components using IClabel → Flag components as artifact menu item.
- Locate component equivalent dipoles using the Tools → Locate dipoles using DIPFIT -> Head model and settings menu item, and then the Tools → Locate dipoles using DIPFIT -> Autofit menu item.
- Extract data epochs using the Tools → Extract epochs menu item.
Filter the datasets
The selected datasets are already processed and do not require additional processing. Nevertheless, we will simply filter them for illustrative purposes. Select the Tools → Filter the data → Basic FIR filter menu item. The following interface pops up. Enter 100 in the second edit box to low-pass filter the data below 100 Hz (this is half the 200 Hz sampling frequency and will not affect subsequent processing). Press Ok.
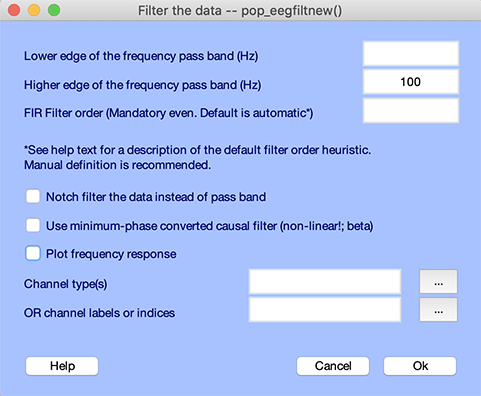
EEGLAB asks for confirmation, warning that the datasets will be automatically overwritten on disk. Select Proceed.
The datasets are filtered one by one and resaved on disk. All menu items mentioned previously work similarly.
Apply ICA
Running ICA on multiple datasets is useful when you have two EEGLAB datasets for two conditions from a subject that were collected in the same session and want to perform ICA decomposition on their combined data. Using this option, you do not have to concatenate the datasets yourself; EEGLAB will detect that these two datasets belong to the same subject, merge them, run ICA, and save the (same) decomposition in each of the subject’s datasets.
The graphic interface for running ICA is a bit more elaborate. Select menu item Tools "> Run ICA. The following window will appear.
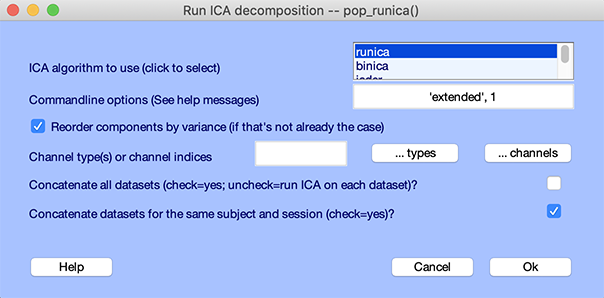
By default, pop_runica.m will concatenate datasets from the same subject and session. For example, you may have several datasets time-locked to different classes of events, constituting several experimental conditions per subject, all collected in the same session with the same electrode montage. By default (leaving the lowest checkbox checked), pop_runica.m will perform ICA decomposition on the concatenated data trials from these datasets, and will then attach the same ICA unmixing weights and sphere matrices to each dataset. Information about the datasets selected for concatenation will be provided on the MATLAB command line before beginning the decomposition. Note that to apply pop_runica.m to concatenated datasets, the datasets’ epoch lengths are assumed to be equal.
If you wish (and have enough computer RAM), you may also ask pop_runica.m to load and concatenate all datasets before running ICA. We do not recommend this approach since it tacitly (and unreasonably) assumes that the very same brain and non-brain sources and the very same electrode positions exist in each session and/or subject.
Multiple-subject selection and EEGLAB studies
After processing of all selected datasets, you may use menu item File → Create Study → Using all loaded datasets to create a study using all loaded datasets (if you only want to use the dataset you selected, you will have to remove the other datasets from the list of datasets to include in the STUDY). See the group analysis tutorial for more details.
Note that you may also create a STUDY before processing datasets and then proceed to batch process them. Once a STUDY is created, all STUDY datasets remain selected, and multi-subject processing menu items become available. See the STUDY creation tutorial on how to create a STUDY.