Extracting data Epochs
Table of contents
Load the sample EEGLAB dataset
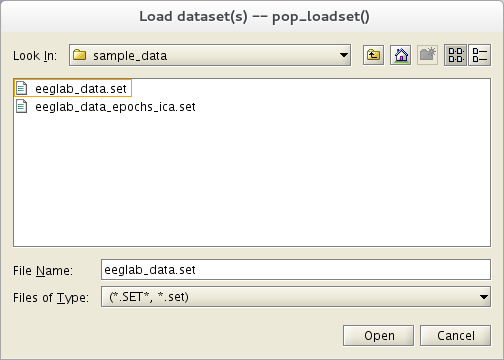
Select the File menu item and press the Load existing dataset sub-menu item. Select the tutorial file “eeglab_data.set” located in the “sample_data” folder of EEGLAB. Then press Open.
Extracting data epochs
To study the event-related EEG dynamics of continuously recorded data, we must extract data epochs time-locked to events of interest (for example, data epochs time-locked to onsets of one class of experimental stimuli) by selecting Tools → Extract Epochs from the EEGLAB main user interface.
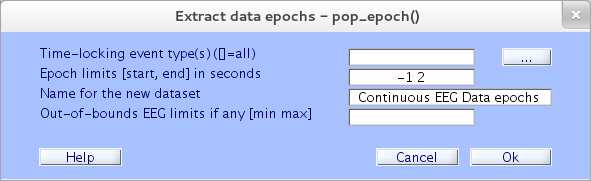
Click on the upper right button, marked “…”, of the resulting pop_epoch.m window, which calls up a browser box listing the available event types.
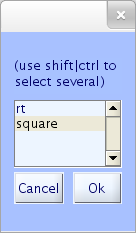
Here, choose event type square (onsets of square target stimuli in this experiment), and press Ok. You may also type in the selected event type directly in the upper text box of the pop_epoch.m window.
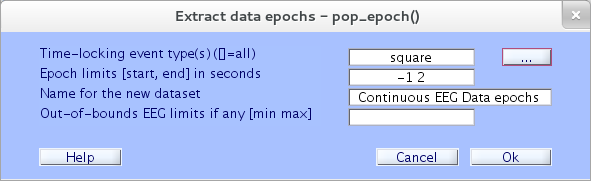
Here, retain the default epoch limits (from 1 sec before to 2 sec after the time-locking event). If you wish, add a descriptive name for the new dataset. Then press Ok. A new window will pop up, offering another chance to change the dataset name and/or save the dataset to a disk file. Accept the defaults and enter Ok.
Another window will then pop up to facilitate the removal of meaningless epoch baseline offsets. This operation is discussed in the next section. In this example, the stimulus-locked windows are 3 seconds long. It is often better to extract long data epochs, as here, to make time-frequency decomposition possible at lower (below 10 Hz) frequencies.
Removing baseline values
Removing a mean baseline value from each epoch is useful when baseline differences between data epochs (e.g., those arising from low-frequency drifts or artifacts) are present. These are not meaningfully interpretable, but if left in the data, could skew the data analysis. However, high-pass filtering data is also a form of baseline correction, rendering the current step optional. It is also essential to remember that baseline correction introduces random offsets in each channel, something ICA and source reconstruction algorithms cannot easily cope with. Baseline correction should thus be used with caution and short baseline windows (on the order of 100 milliseconds) avoided when possible.
After the data has been epoched, the following window will pop up automatically. It is also possible to call it directly by selecting the Tools → Remove baseline menu item.
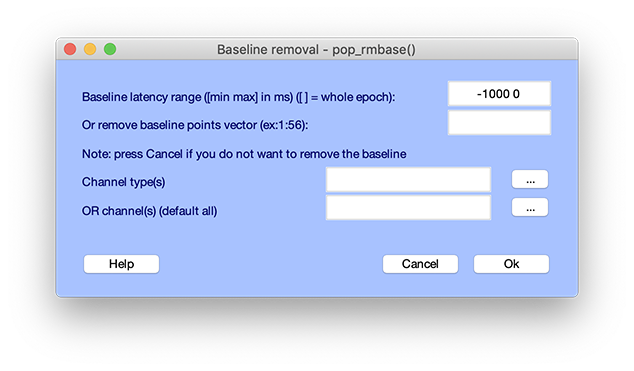
Here we may specify the baseline period in each epoch (in ms) – the latency window in each epoch across which to compute the mean to remove The original epoched dataset is by default overwritten by the baseline-removed dataset. Note: There is no uniformly ‘optimum’ method for selecting either the baseline period or the baseline value. Using the mean value in the pre-stimulus period (the pop_rmbase.m default) is effective for many datasets if the goal of the analysis is to define transformations that occur in the data following the time-locking events. By default, baseline removal will be applied to all channels. However, you can also choose specific channels by type (can be specified while editing channel information), or manually select them. Click on the ‘…’ push buttons to see the list of available types/channels for selection. Press Ok to subtract the baseline (or Cancel to not remove the baseline).
A new window will pop up to change the dataset name and/or save the dataset to a disk file. Accept the defaults and enter Ok.
Extracting sub-epochs
Even after data epochs have been extracted, it is possible to extract sub-epochs with a reduced time range. This is done using the function pop_select.m called by selecting Edit → Select data. The example below would select data sub-epochs with the epoch time range from -500 ms to 1000 ms.
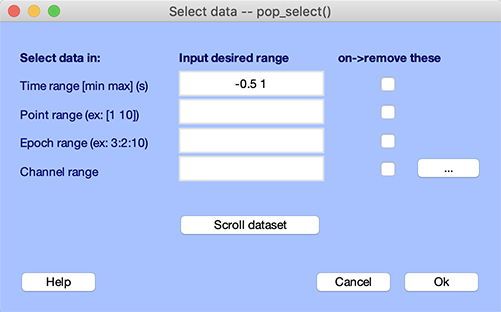
Selecting data epochs
There is no real good reason to select subsets of data epochs. When comparing conditions – performed by creating contrast at the STUDY level (the group analysis interface which may also be used for single-subject analysis) – one may ignore specific data epochs.
Nevertheless, they may be cases in which you might want to remove specific artifactual or irrelevant data epochs.
Selecting a subset of data epochs
Select menu item File and press sub-menu item Load existing dataset. Select the tutorial file “eeglab_data_epochs_ica.set” located in the “sample_data” folder of EEGLAB. Then press Open.
The simplest way to remove data epochs is by selecting epoch indices. This is done using the function pop_select.m called by selecting Edit → Select data. The example below select epochs 1 to 10 (1:10 is MATLAB notation to indicate 1, 2, 3, 4, 5, 6, 7, 8, 9, 10).
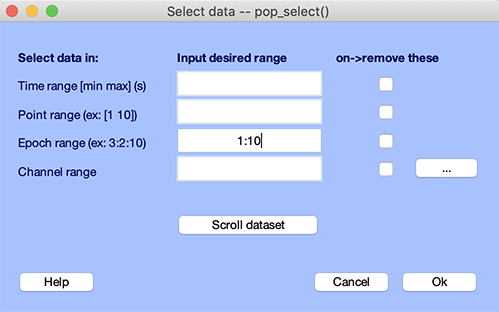
Alternatively, epochs 1 to 10 may be removed by checking the checkbox adjacent to the epoch edit box.
Selecting a subset of data epochs based on event information
In the EEGLAB sample dataset, half the targets appeared at position 1 and the other half at position 2. In this section, we will keep all position 1 data epochs. Import the “eeglab_data_epochs_ica.set” tutorial dataset, as indicated in the previous section.
Select Edit → Select epochs or events. The pop_selectevent.m window (below) will appear. Enter “1” in the textbox next to position, which will select all epochs in which the target appeared in position 1 (note the selected checkbox Remove epochs not referenced by any selected event). Press Ok.
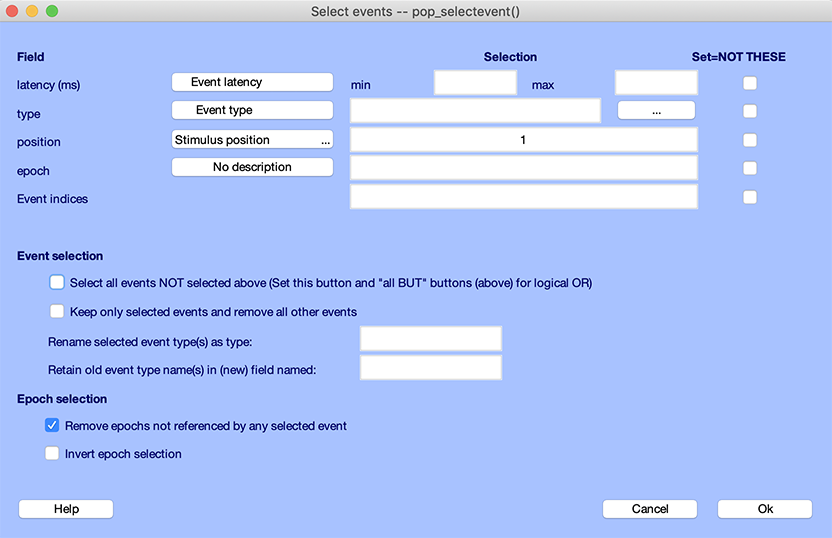
Note: The option set above the Cancel button (above) Remove epochs not referenced by any selected event. If this checkbox were left unset and the checkbox Keep only selected events and remove all other events, the function would select the specified events but would not remove epochs not containing those events.
The confirmation window below appears. Press Ok.
Now a pop_newset.m window for saving the new dataset pops up. We name this new dataset “Square, Position 1” and press Ok.