Plotting channel spectra and maps
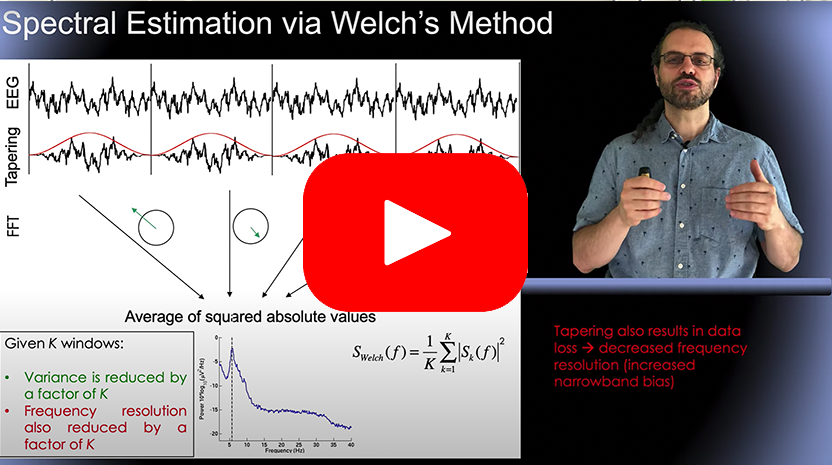
Besides reading the tutorial sections below, you may want to watch the short video on computing spectra in EEGLAB (hosted on Youtube) below. In particular, we recommend video 1 and 2 describing the Welch method used in this section, and video 5, describing the EEGLAB functions used in this section.
Load the sample EEGLAB dataset
Select the File menu item and press the Load existing dataset sub-menu item. Select the tutorial file “eeglab_data.set” located in the “sample_data” folder of EEGLAB. Then press Open.
Plot Channel Spectra and Maps
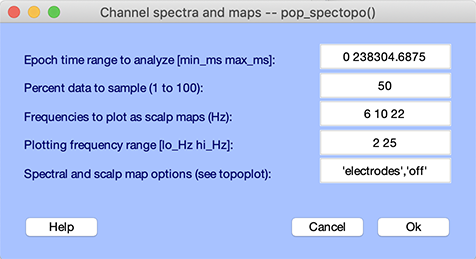
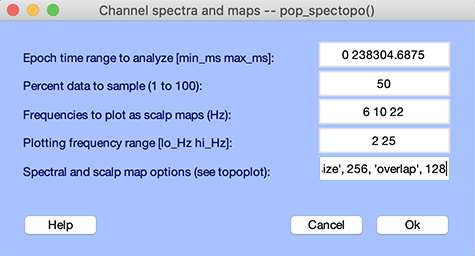
To plot the channel spectra and associated topographical maps, select Plot → Channel spectra and maps. This will pop up the pop_spectopo.m window (below). Leave the default settings and press Ok.
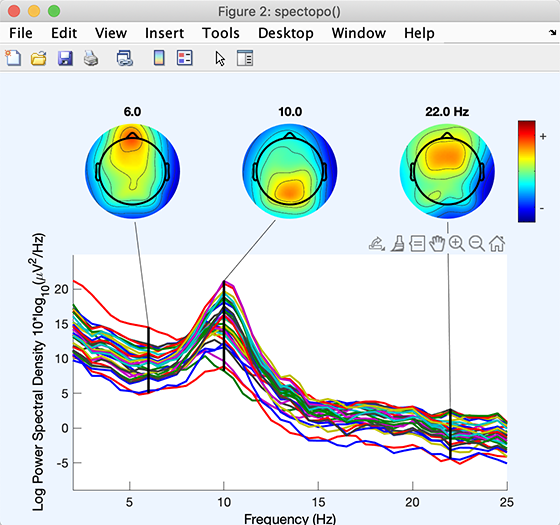
The function should return a spectopo.m plot (below). Since we only sampled 50% of the data (via the Percent data… edit box above), results should differ slightly on each call. (Naturally, this will not occur if you enter 100% in the edit box).
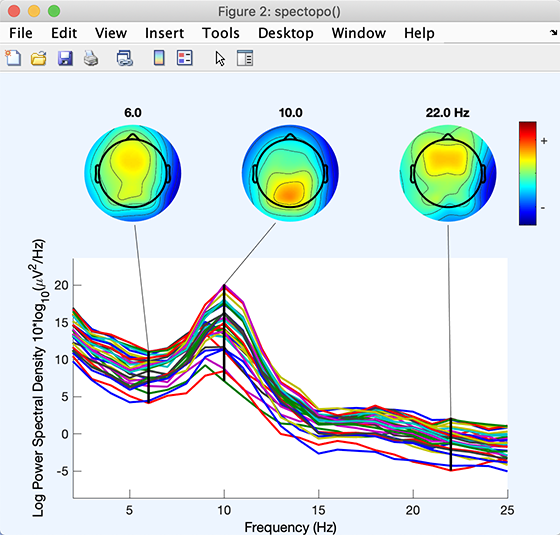
Each colored trace represents the spectrum of the activity of one data channel. The leftmost scalp map shows the scalp distribution of power at 6 Hz, which in these data is concentrated on the frontal midline. The other scalp maps indicate the distribution of power at 10 Hz and 22 Hz.
The pop_spectopo.m window menu (above) allows the user to compute and plot spectra in specific time windows in the data. The Percent data… value can be used to speed the computation (by entering a number close to 0) or to return more definitive measures (by entering a number closer to 100).
On the MATLAB command line, the parameters for calculating the spectrum using the Welch method are exposed (window size of 128 samples with no overlap between windows). We can change these parameters. Select menu item Plot → Channel spectra and maps and in the Spectral and scalp map options edit box, enter “ ‘winsize’, 256, ‘overlap’, 128 “.
This will result in a smoother spectrum with higher frequency resolution as shown below.
Note that functions pop_spectopo.m and spectopo.m also work with epoched data.
Another menu item, Plot → Channel properties, plots the scalp location of a selected channel, its activity spectrum, and an ERP-image plot of its activity in single-epochs.
Note: The MATLAB Signal Processing Toolbox should be in your MATLAB path to use these functions. EEGLAB has replacement functions in case the signal processing toolbox is not present, but their capabilities are limited.
Note: It is also possible to plot electrode locations in the spectral graph by entering ‘’ ‘electrodes’, ‘on’ ‘’ in the lowest text box (Scalp map options) of the interactive pop_spectopo.m window.